如何将echarts图例与曲线显示顺序不一致
Posted
tags:
篇首语:本文由小常识网(cha138.com)小编为大家整理,主要介绍了如何将echarts图例与曲线显示顺序不一致相关的知识,希望对你有一定的参考价值。
Excel中图表的图例的默认顺序为数据系列的添加顺序,如果后期想修改图例顺序,方法是:右键菜单→添加数据→在“选择源数据”对话框中调整数据系列的顺序。
下面以Excel 2010为例演示此过程:
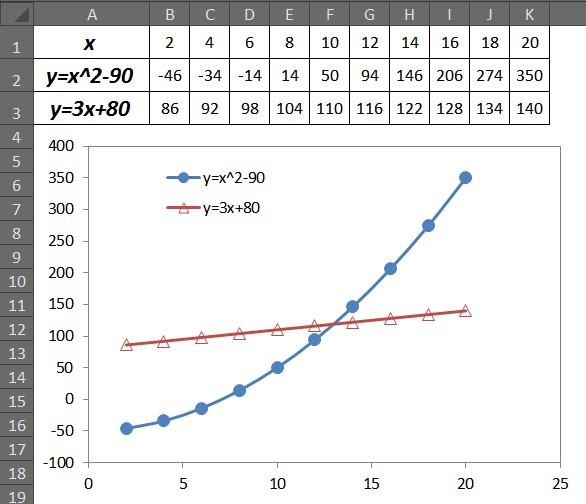
1、默认的源数据及图例顺序:从图中可知默认的图例顺序与表格中源数据顺序一致
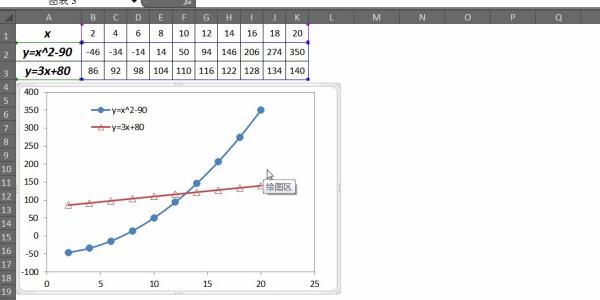
2、点击图片查看修改图例顺序的动画演示:
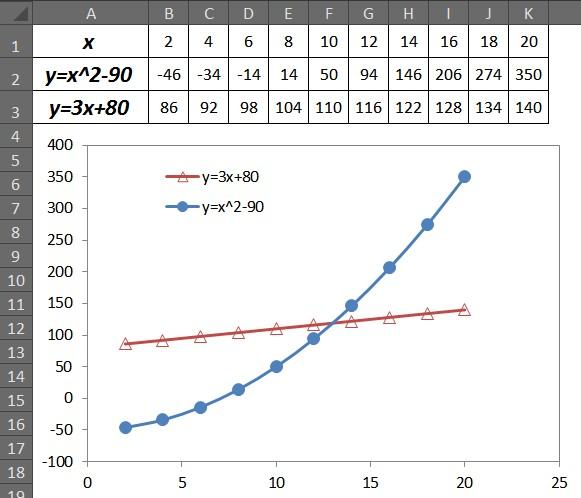
3、最终效果如下:源数据顺序未变而图例顺序已经被调整
同求echarts的解决方法 参考技术B 在狱咏蝉(骆宾王)
反转图例的顺序
【中文标题】反转图例的顺序【英文标题】:Invert the order of the legend 【发布时间】:2021-10-18 21:22:49 【问题描述】:我已经编写了一个函数来为我闪亮的仪表板创建曲线图。传说的顺序似乎不对。出于某种原因,它显示了数据框中的内容的反向。我希望将“其他”类别放在图例的最后。有没有办法改变图例的排序?
plot_maker <- function(df, var, legend_pos = -0.5, legend = TRUE)
temp <- df %>%
pivot_wider(names_from = Year_month, values_from = count) %>%
ungroup() %>%
mutate_at(vars(contains("20")), function(x) x / sum(x, na.rm = TRUE)) %>%
pivot_longer(!(!!sym(var)), names_to = "Year_month", values_to = "value")
temp <- temp %>% inner_join(df %>%
mutate(Year_month = as.character(Year_month)),
by = c(var, "Year_month")
)
temp$Year_month <- factor(temp$Year_month,
levels = c(unique(temp$Year_month))
)
if (sum(temp$value, na.rm = TRUE) == 0)
return(plot.new())
else
p <- plot_ly(
data = temp,
y = ~value,
x = ~Year_month,
color = ~ (get(var)),
legendgroup = ~ (get(var)),
hoverinfo = "text",
text = ~ paste(
get(var),
"<br>Count:", count,
"<br>PCT:", sprintf("%1.2f%%", 100 * value)
)
) %>%
add_lines() %>%
layout(
yaxis = list(
tickformat = "%",
title = ""
),
xaxis = list(title = ""),
legend = list(
orientation = "h", yanchor = "bottom", y = legend_pos,
font = list(size = 10),
traceorder= 'normal'
),
template = "plotly_dark"
)
if (legend == FALSE)
p <- p %>% layout(showlegend = FALSE)
return(p)
运行函数
plot_maker(df1, "therapy_class")
这是测试数据(df1)
structure(list(therapy_class = structure(c(1L, 1L, 1L, 1L, 1L,
1L, 2L, 2L, 2L, 2L, 2L, 2L, 3L, 3L, 3L, 3L, 3L, 3L, 4L, 4L, 5L,
5L, 5L, 5L, 5L, 5L, 6L, 6L, 6L, 6L, 6L, 6L, 7L, 7L, 7L, 7L, 7L,
7L, 8L, 8L, 8L, 8L, 8L, 8L, 9L, 9L, 9L, 9L, 9L, 9L), .Label = c("ALK Inhibitors",
"Anti-VEGF-based therapies", "EGFR TKIs", "EGFR-antibody based therapies",
"Non-platinum-based chemotherapy combinations", "IO-based therapies",
"Platinum-based chemotherapy combinations", "Single agent chemotherapies",
"Other"), class = c("ordered", "factor")), Year_month = structure(c(2020.91666666667,
2021, 2021.08333333333, 2021.16666666667, 2021.25, 2021.33333333333,
2020.91666666667, 2021, 2021.08333333333, 2021.16666666667, 2021.25,
2021.33333333333, 2020.91666666667, 2021, 2021.08333333333, 2021.16666666667,
2021.25, 2021.33333333333, 2021.08333333333, 2021.16666666667,
2020.91666666667, 2021, 2021.08333333333, 2021.16666666667, 2021.25,
2021.33333333333, 2020.91666666667, 2021, 2021.08333333333, 2021.16666666667,
2021.25, 2021.33333333333, 2020.91666666667, 2021, 2021.08333333333,
2021.16666666667, 2021.25, 2021.33333333333, 2020.91666666667,
2021, 2021.08333333333, 2021.16666666667, 2021.25, 2021.33333333333,
2020.91666666667, 2021, 2021.08333333333, 2021.16666666667, 2021.25,
2021.33333333333), class = "yearmon"), count = c(18L, 17L, 16L,
15L, 15L, 15L, 37L, 39L, 38L, 47L, 48L, 32L, 63L, 45L, 64L, 73L,
63L, 57L, 1L, 1L, 6L, 5L, 5L, 12L, 2L, 8L, 312L, 327L, 296L,
371L, 324L, 307L, 127L, 115L, 99L, 148L, 124L, 141L, 69L, 51L,
48L, 66L, 58L, 38L, 45L, 44L, 34L, 43L, 64L, 52L)), row.names = c(NA,
-50L), groups = structure(list(therapy_class = structure(1:9, .Label = c("ALK Inhibitors",
"Anti-VEGF-based therapies", "EGFR TKIs", "EGFR-antibody based therapies",
"Non-platinum-based chemotherapy combinations", "IO-based therapies",
"Platinum-based chemotherapy combinations", "Single agent chemotherapies",
"Other"), class = c("ordered", "factor")), .rows = structure(list(
1:6, 7:12, 13:18, 19:20, 21:26, 27:32, 33:38, 39:44, 45:50), ptype = integer(0), class = c("vctrs_list_of",
"vctrs_vctr", "list"))), row.names = c(NA, -9L), class = c("tbl_df",
"tbl", "data.frame"), .drop = TRUE), class = c("grouped_df",
"tbl_df", "tbl", "data.frame"))
【问题讨论】:
【参考方案1】:p <- plot_ly(
data = temp,
y = ~value,
x = ~Year_month,
color = ~ (get(var)),
legendgroup = ~ (get(var)),
hoverinfo = "text",
text = ~ paste(
get(var),
"<br>Count:", count,
"<br>PCT:", sprintf("%1.2f%%", 100 * value)
)
) %>%
add_lines() %>%
layout(
yaxis = list(
tickformat = "%",
title = ""
),
xaxis = list(title = ""),
legend = list(
orientation = "h", yanchor = "bottom", y = legend_pos,
font = list(size = 10),
traceorder= 'reversed'
),
template = "plotly_dark"
)
【讨论】:
以上是关于如何将echarts图例与曲线显示顺序不一致的主要内容,如果未能解决你的问题,请参考以下文章
当数据中不存在分组变量的所有级别时,绘图之间的颜色比例和图例一致